|
nmo.gather
Figure 2 A synthetic shot gather. The first and tenth trace were selected to test the algorithm. | 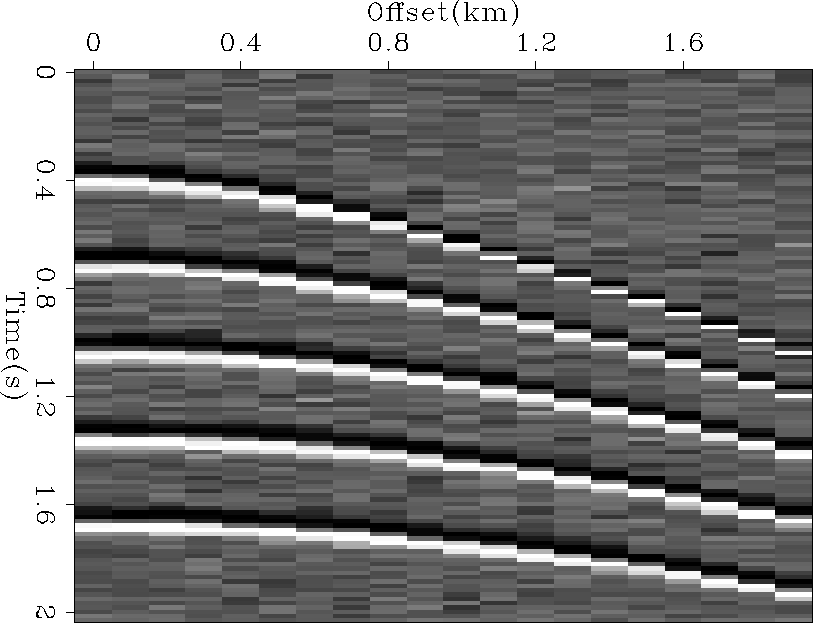 |
![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) ).
We selected two traces some distance apart (the first and tenth trace) and applied
the algorithm. The two traces can be seen in the left part of Figure
).
We selected two traces some distance apart (the first and tenth trace) and applied
the algorithm. The two traces can be seen in the left part of Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) .
Note the time-variant alignment error.
.
Note the time-variant alignment error.
|
nmo.gather
Figure 2 A synthetic shot gather. The first and tenth trace were selected to test the algorithm. |  |
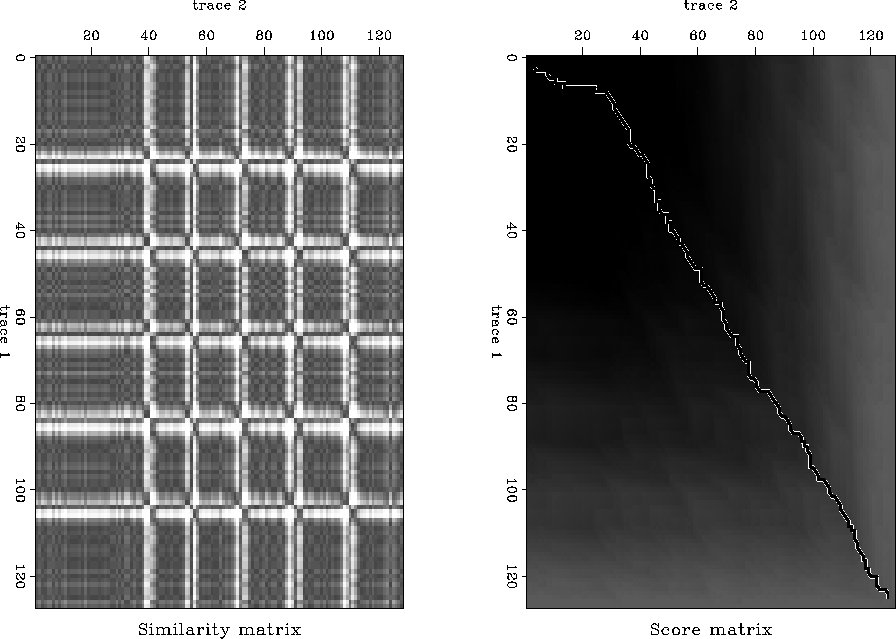
Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) shows the similarity (left) and score (right) matrices.
The black lines in the similarity matrix represent low scores and correspond to the events in the data, they only
disappear (or match) when encountering another wavelet. The score matrix
shows exactly what we expect to see. A slightly non-diagonal maximum (except
for edge effects at low times corresponding to a lack of coherent events).
Figure
shows the similarity (left) and score (right) matrices.
The black lines in the similarity matrix represent low scores and correspond to the events in the data, they only
disappear (or match) when encountering another wavelet. The score matrix
shows exactly what we expect to see. A slightly non-diagonal maximum (except
for edge effects at low times corresponding to a lack of coherent events).
Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) shows the input (left) and output (right) along
with their corresponding differences. The output is much better aligned and
the overall differences reduced.
The difference trace is a proxy for guaging the quality of alignment, but the goal is
not to drive this difference to zero. The algorithm keys on strong events whose
alignment may result in sizable differences at other levels. This is a significant
departure from () who minimize a difference
measure to determine alignment.
shows the input (left) and output (right) along
with their corresponding differences. The output is much better aligned and
the overall differences reduced.
The difference trace is a proxy for guaging the quality of alignment, but the goal is
not to drive this difference to zero. The algorithm keys on strong events whose
alignment may result in sizable differences at other levels. This is a significant
departure from () who minimize a difference
measure to determine alignment.
|
nmo.in-out
Figure 3 The left plot shows the two input traces and the right plot the traces after alignment. The third trace in each display shows the difference. | 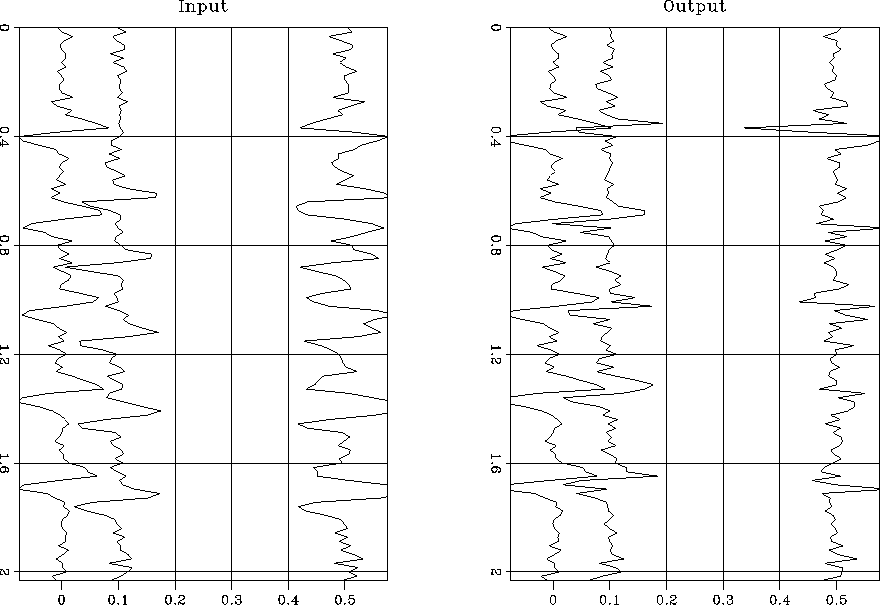 |
 |
![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) and the left panel
is the similarity matrix. Axes labels refer to time sample numbers (not seconds).
and the left panel
is the similarity matrix. Axes labels refer to time sample numbers (not seconds).
For a second test we chose a common reflection point (CRP) gather
from a 2-D marine
dataset (Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) ).
The gather is an angle gather (, ) after
phase-shift plus-interpolation (PSPI) migration (). Note that
we still see some residual moveout in the angle gather.
The left panel of Figure
).
The gather is an angle gather (, ) after
phase-shift plus-interpolation (PSPI) migration (). Note that
we still see some residual moveout in the angle gather.
The left panel of Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) shows the input two traces (third
and sixteenth).
shows the input two traces (third
and sixteenth).
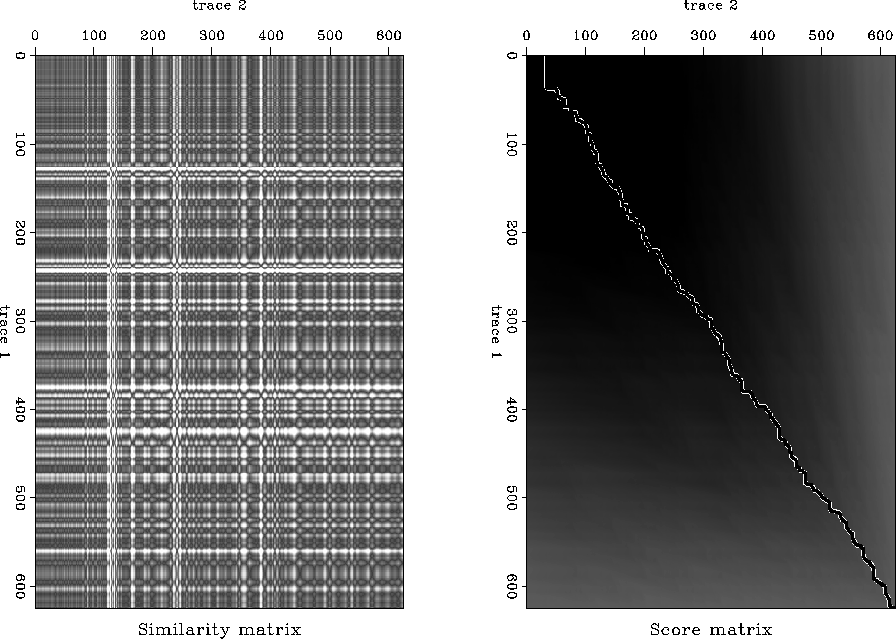
After running the algorithm we obtained the score and similarity matrices
seen in Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) . Note how the structure of
the similarity matrix to the previous example (Figure
. Note how the structure of
the similarity matrix to the previous example (Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) ).
The score matrix and the corresponding maximum has the shape that we would anticipate.
It is generally diagonal with some deviations. The output two traces appear to be
better aligned (the right panel of Figure
).
The score matrix and the corresponding maximum has the shape that we would anticipate.
It is generally diagonal with some deviations. The output two traces appear to be
better aligned (the right panel of Figure ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) ), but the
difference isn't as reduced as we would hope. Our belief is this caused
by a poor stretching algorithm.
), but the
difference isn't as reduced as we would hope. Our belief is this caused
by a poor stretching algorithm.
|
big.gather
Figure 5 The CRP gather used for trace alignment. The third and 16th trace were used. |  |
 |
|
big.in-out
Figure 7 The left plot shows the two input traces and the right plot the traces after alignment in the window from three to four seconds. The third trace in each display shows the difference. | 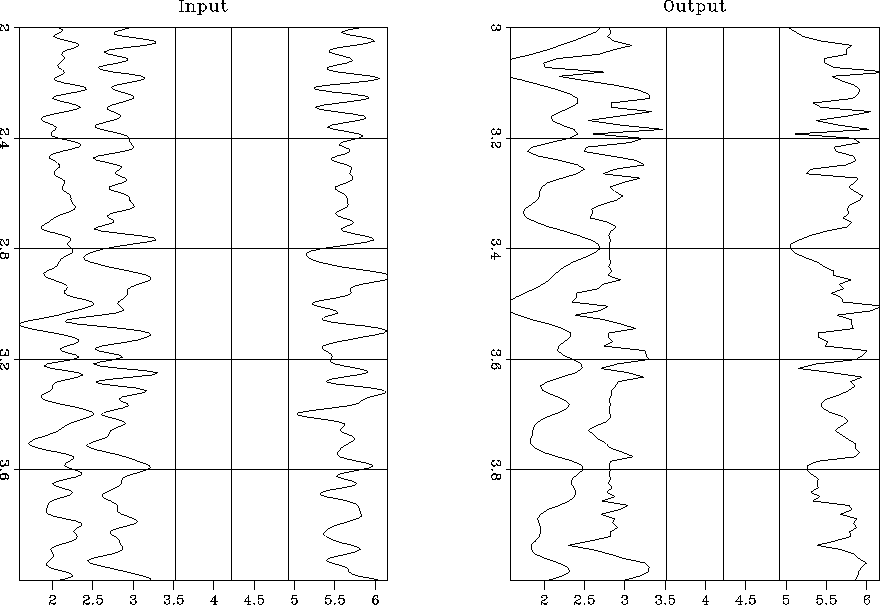 |